Chromatin, the intricate combination of DNA and histone proteins, plays a pivotal role in governing nuclear processes. In this article, our focus is on the dynamic SWI/SNF chromatin remodeling complex. Essential to the epigenetic regulation of numerous nuclear processes, ranging from transcription to DNA repair and replication, SWI/SNF stands as a key player in the complex world of chromatin dynamics.
Chromatin Structure and Dynamics: A Brief Overview
Nucleosomes, the basic units of chromatin, create a barrier to protein access within the genome. However, these nucleosomes are not static; their occupancy and positioning are subject to alteration by chromatin remodeling complexes (CRCs). Using energy derived from ATP hydrolysis, CRCs, including the SWI/SNF complex, disrupt electrostatic interactions between histones and DNA.
SWI/SNF: A Unique Chromatin Remodeling Player
Among the four main classes of CRCs found in yeast, animals, and plants (Imitation Switch, SWI; Inositol-Requiring 80, INO80; Chromodomain Helicase DNA-Binding, CHD; and Mating Type Switch Deficient Sucrose Nonfermenting, SWI/SNF), the SWI/SNF complex stands out. Its distinct ability to modify the accessibility of genomic DNA within chromatin sets it apart. SWI/SNF achieves this by not only sliding the histone octamer along the DNA but also by ejecting it, influencing crucial nuclear processes like transcription, DNA repair, and replication.
SWI/SNF Architecture: Modules and Functions
SWI/SNF CRCs are colossal protein assemblies, weighing between 1 and 2 megadaltons. Comprising multiple conserved and interconnected subunits, including a central ATPase, these complexes are organized into three modules: the motor module, the actin-related protein (ARP) module, and the base module.
- Motor Module: This powerhouse hydrolyzes ATP and disrupts DNA-histone contacts, playing a pivotal role in reshaping chromatin structure.
- ARP Module: Serving as a connector between the motor and base modules, ARP enhances the coupling of ATP hydrolysis to DNA translocation, ensuring efficient chromatin remodeling.
- Base Module: Engaging with histones, this module’s interactions are influenced by histone posttranslational modifications and specific DNA sequence features, adding another layer of complexity to the chromatin remodeling process.
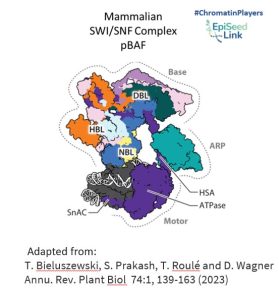
Activities of SWI/SNF Complexes in the Nucleus
Within the nucleus, SWI/SNF remodelers orchestrate transcriptional regulation at various levels. They participate in modulating enhancer activity, recruiting the general transcriptional machinery, and facilitating the transition from paused to elongating RNA polymerase.
SWI/SNF is recruited to genomic DNA by transcription factors (TFs), unable to bind DNA directly. Pioneering TFs uniquely access DNA wrapped in nucleosomes, recruiting SWI/SNF to reduce nucleosome occupancy or move nucleosomes away from enhancers. This process enables the binding of additional TFs required for complete transcriptional output.
Additionally, SWI/SNF facilitates the access of RNA polymerase II to the promoter, contributing to the initiation of transcription. Notably, SWI/SNF complexes push the +1 nucleosome away from the transcription start site (TSS), enabling the recruitment of RNA polymerase II and the general transcriptional machinery.
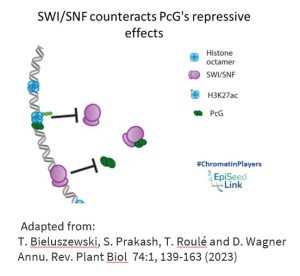

SWI/SNF in Epigenetic Regulation: Interplay with Polycomb Group Proteins
Polycomb group (PcG) epigenetic regulators, known for silencing transcriptional programs, intersect with SWI/SNF in a fascinating manner. PcG complexes PRC2 and PRC1 deposit repressive histone modifications, including H3K27me3. SWI/SNF CRCs, acting as molecular gatekeepers, eject PcG complexes and impede their access to chromatin at numerous loci. This dynamic interplay between PcG and SWI/SNF contributes to shaping the accessible chromatin landscape during developmental processes in both animals and plants.
Unraveling SWI/SNF Complexity in Plants: Insights from Arabidopsis
In the plant kingdom, understanding the classification and composition of SWI/SNF complexes has been a challenge. In Arabidopsis thaliana three types of SWI/SNF ATPase has been identified: BRAHMA (BRM/CHR2), SPLAYED (SYD/CHR3), MINUSCULE1 (MINU1/CHR12), and its closed related MINUSCULE2 (MINU2/CHR23). These ATPases, though conserved in the ATPase domain, exhibit diverse domain structures at the N-terminal and C-terminal regions, suggesting functional divergence. They also form different complexes (BAS, SAS or MAS) by interacting with different groups of common or specific subunits.
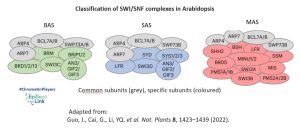

Mutant analysis of BRM, SYD, MINU1, and MINU2 offers a glimpse into the crucial role these ATPases play in plant development. Single mutants, such as brm and syd, exhibit pleiotropic defects—reduced plant size, slow growth, curled rosette leaves, aberrant flower development, and sterility. BRM and SYD occupy thousands of common target genes and exhibit similar binding patterns, indicating that the two enzymes are functionally related.
The brm syd double mutant faces embryonic lethality, emphasizing the essential nature of these ATPases in early embryo development.
Loss-of-function minu1 and minu2 single mutants are morphologically indistinguishable from the wild type, while the minu1/2 (minu1 and minu2) double mutant exhibits embryonic lethal phenotypes. The weak minu1/2 double mutant forms small and bushy plants that have defects in maintenance of both root and shoot apical meristems. These results suggest that the Arabidopsis SWI/SNF ATPases have specific functions in regulating development, although the mechanisms underlying the specificity are mainly unknown.
Analyzing the phenotype of the mutants of three types of SWI/SNF complex concluded that three classes of SWI/ SNF complexes have both overlapping and specific functions in the regulation of various developmental processes, including the development of leaves, gametophytes, embryos and flowers.
Moreover, it has been shown that the three types of SWI/SNF complexes differently regulate chromatin accessibility, providing a plausible explanation for the specific functions of different SWI/SNF complexes.
Analysis of mutants representing three types of SWI/SNF complexes revealed overlapping and specific functions in the regulation of diverse developmental processes, encompassing leaf, gametophyte, embryo, and flower development. Additionally, distinct regulatory roles were observed as the three SWI/SNF complex types differentially influenced chromatin accessibility, providing insight into their specific functionalities.
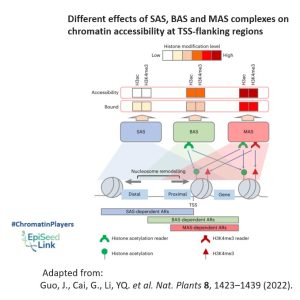
Examining genomic distribution patterns, all three plant SWI/SNF complexes predominantly bind to the transcription start site (TSS)-flanking regions of protein-coding genes, indicating shared intrinsic features facilitating this binding. However, the BAS complex primarily targets the TSS-flanking region, while SAS and MAS extend their distribution to the upstream distal and downstream intragenic regions, respectively. This divergence is suspected to arise from varying binding affinities for active histone modifications H3ac and H3K4me3, which are enriched at the intragenic region.
The distinct binding abilities of BAS, SAS, and MAS for active histone modifications likely dictate their selective binding to different gene regions. Consequently, active histone modifications regulate the function of these complexes at both the chromatin-binding and nucleosome remodeling levels, although the precise molecular mechanisms remain elusive.
The proposed mechanism involves specific modified-histone readers within SWI/SNF complexes, responsible for recognizing histone modifications and determining the diverse regulation of nucleosome remodeling activities across the three SWI/SNF complex classes. This interplay between complex composition and histone modification patterns highlights the evolution of SWI/SNF complexes with a flexible selection of distinct histone readers, showcasing their fine-tuned adaptation to transcriptional regulation in Arabidopsis.
In conclusion, this review highlights the vital role of the SWI/SNF chromatin remodeling complex in Arabidopsis development, shedding light on the distinct functions of its ATPase subunits. As we explore chromatin accessibility and genomic distribution patterns, it becomes evident that further research is essential to uncover the intricate mechanisms governing SWI/SNF-mediated transcriptional regulation in plants.
References
- Bieluszewski, T., Prakash, S., Roulé, T., Doris Wagner, D. The Role and Activity of SWI/SNF Chromatin Remodelers. Annu. Rev. Plant Biol 74:1, 139-163 (2023) https://doi.org/10.1146/annurev-arplant-102820-093218
- Guo, J., Cai, G., Li, YQ. et al. Comprehensive characterization of three classes of Arabidopsis SWI/SNF chromatin remodeling complexes. Nat. Plants 8, 1423–1439 (2022). https://doi.org/10.1038/s41477-022-01282-z
Text by Carlos Gámez Álvarez, PhD Student EpiSeedLink Marie Skłodowska-Curie Actions
Frontal image from https://bit.ly/3RrxEP7

